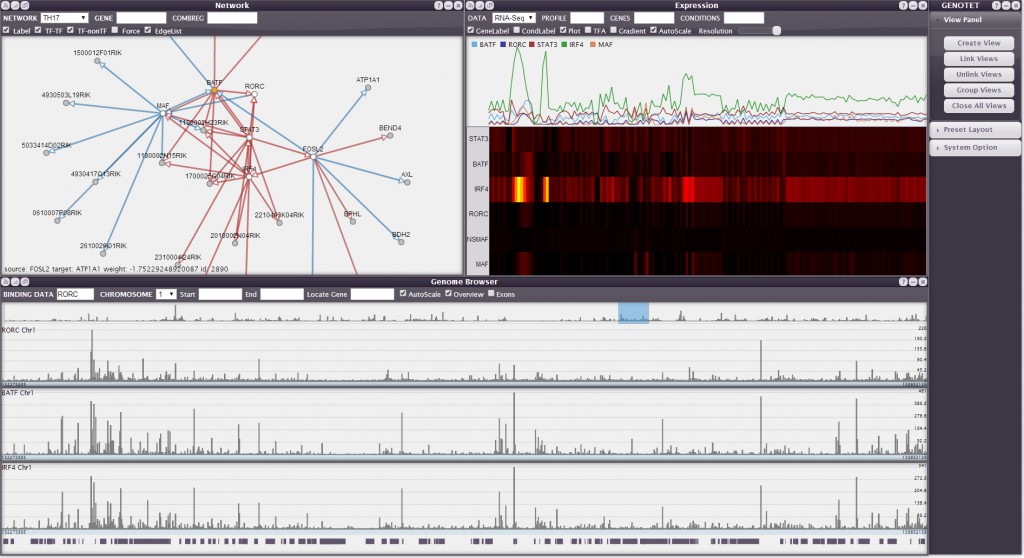
Transcription factors (TFs) are proteins that bind to gene promoters and enhancers to alter gene expression patterns. Biological regulatory networks are composed of large numbers of such regulatory interactions. Understanding these networks is a fundamental goal in biology. In this work we present an efficient querying model by designing the system Genotet, to support interactive analysis of the gene regulatory network data using specialized data structures and multiple coordinated views. We show the effectiveness of our framework through case studies for the mouse immune system and a model bacteria.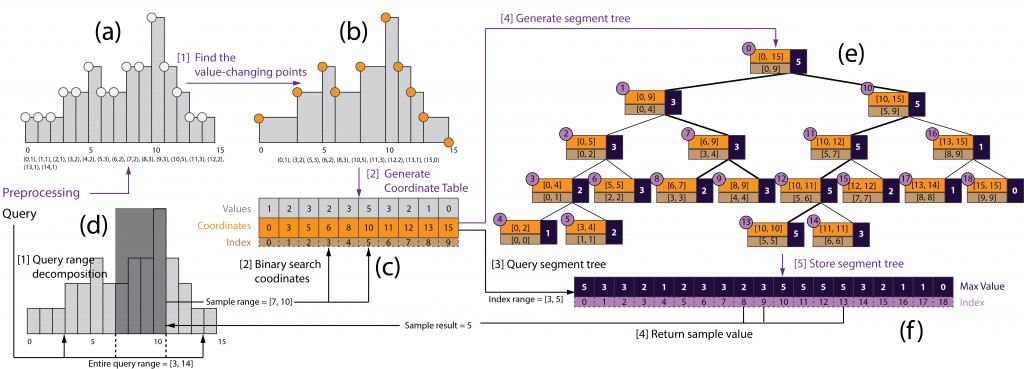
![]()
![]() Large data handling scheme of Genotet. Redundancies were first removed. A custom coordinate table and a flattened segment tree array were built for each binding track in order to support efficient query on the binding data.
Large data handling scheme of Genotet. Redundancies were first removed. A custom coordinate table and a flattened segment tree array were built for each binding track in order to support efficient query on the binding data.
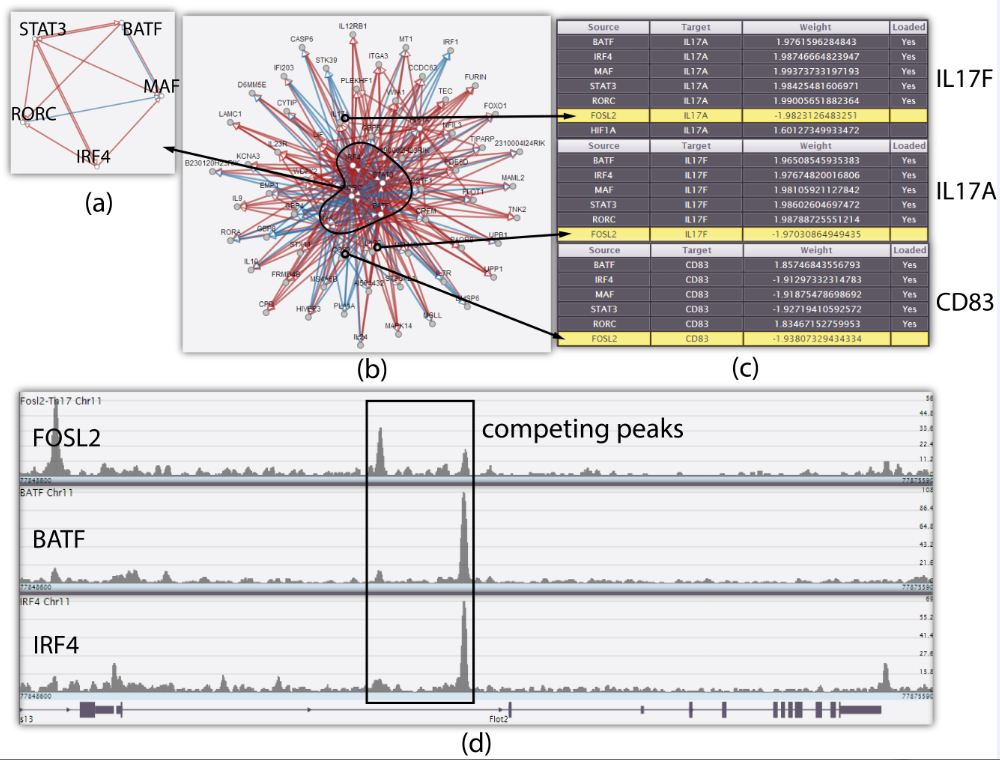
Using Genotet, the biologist was able to validate the finding from a biology journal paper. In the case study, the biologist located the competing peak of a newly found regulator FOSL2.