By Team Glosy: Hong Gao, Mingsi Long, Yulin Shen, and Jie Yang
Objectives
Osteoarthritis (OA) is a synovial joint tissues disease which lead to pain and long term disabilities. Overtime, patients may develop sufficient progression such that they are eligible for total knee replacement (TKR). Knowing this ahead of time can help clinicians and researchers to start early intervention including clinical trials to the right patients. One way is to use deep learning in Magnetic Resonance Imaging (MRI) to learn representations that signal the TKR. Goal: Use 3D MRI to predict the probability of TKR within eight years.
Example of Data Used
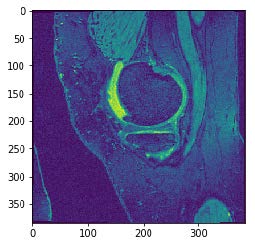
Data Sources & Cohorts
Data Source: NIH Osteoarthritis Initiative (OAI)
- Multicenter, prospective, longitudinal
- 4,796 participants ages 45-79 from 2004
- Clinical information including 3D MRI
- About 10% of the patients received TKR
Supervised Cohort
There are 1,170 patients in the supervised cohort. We define cases as individuals with a positive OA progression outcome in either knee after the baseline enrollment date; individuals who receive a partial knee replacement in either knee or have a knee replacement before baseline are removed. We define controls as individuals who appear at the 96-month visit and do not have a positive OA progression outcome in either knee by that visit.
Unsupervised Cohort
The subjects who are not selected for the supervised cohort will form unsupervised cohort. This cohort includes 3,626 subjects.
Results: Supervised
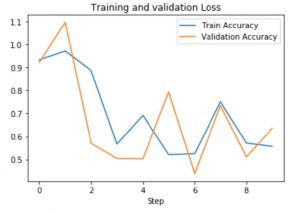
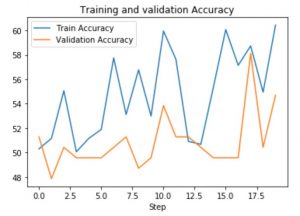
Results: Semi-Supervised Learning
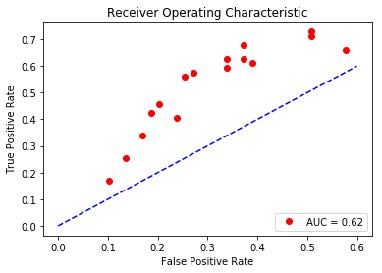
Method 1
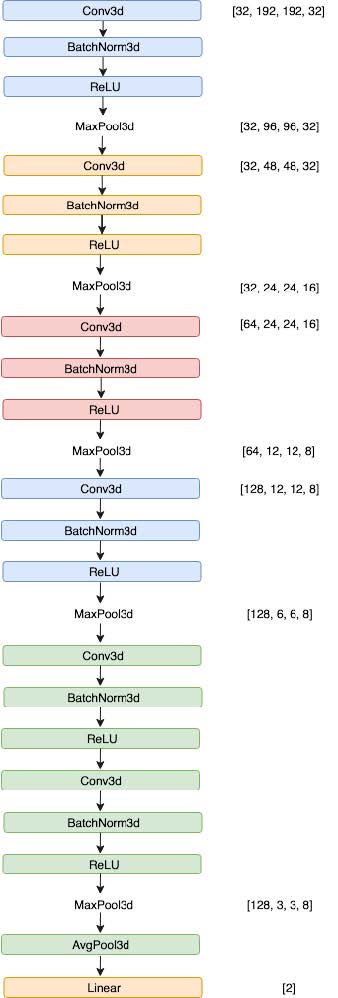
Our supervised learning model GlosyNet is based on the Very
Deep Convolutional Networks (VGG). It took 1,170 fixed-size
448 × 448 × 160 3 dimensional MRI images, cropped them to size of 348 × 348 × 32 combined with various augmentation mechanisms including randomly crop and flip. Processed images are passed through a stack of convolutional (conv.) layers, where we used 3 × 3 filters (Figure 5).
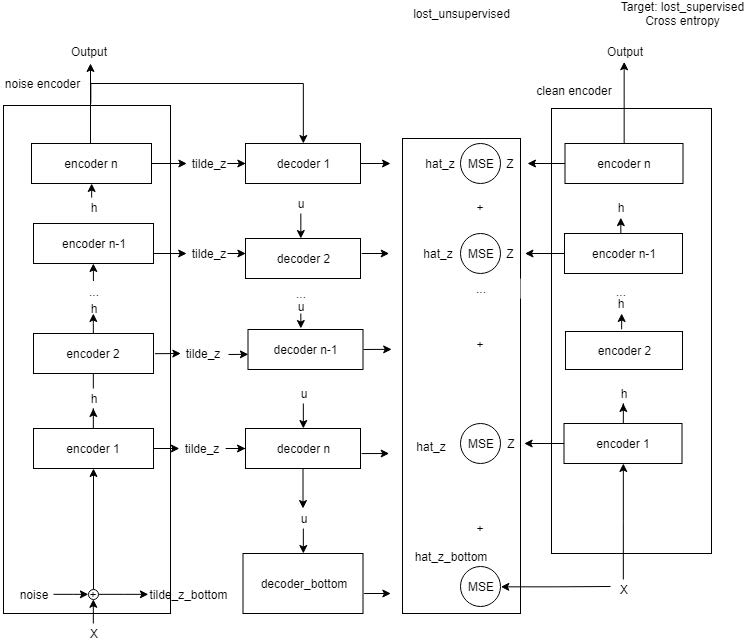
Method 2
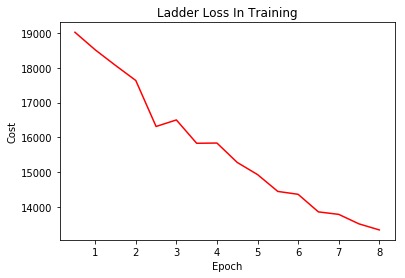
We used Ladder Network, a semi-supervised network. It make use of both labeled and unlabeled images for training. We used GlosyNet as the encoder and generated the cross entropy error. The unsupervised loss is the mean square error of the original image representation and the reconstructed image representation with noise from the decoder.
Recommended Future Steps
- Incorporate patient demographics and clinical information, such as pain level
- Tune hyperparameter of both GlosyNet and the LadderNet based on GlosyNet
- Take advantage of more data collected from the dataset
Sponsor
Sponsor: Cem Deniz, PhD
Department NYU Department of Radiology Web
References
- Rasmus A. et al. Semi-supervised learning with ladder networks. MIT Press, 2015.
- Simonyan K. Zisserman A. Very deep convolutional networks for large-scale image recognition. arXiv, page 1409.1556, 2014.
